  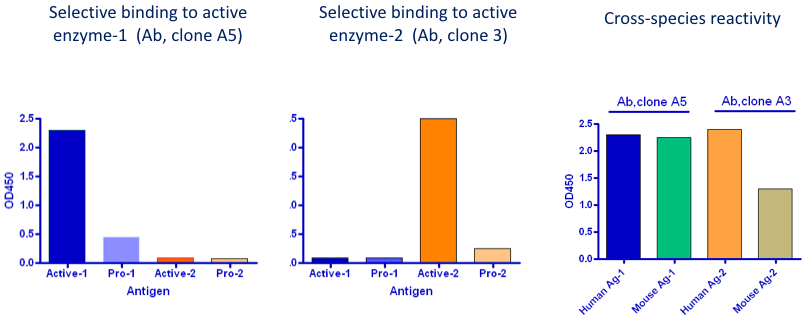
Misincorporation mapping takes advantage of the ability of m 1A and numerous other modified nucleotides to induce misincorporations during the reverse transcription step common to most RNA-Seq protocols. Here, to address the question of the prevalence and location of m 1A in the transcriptome, we used both a high-resolution m 1A-mapping method as well as a bioinformatic approach, termed “misincorporation mapping”. Additionally, whether m 1A sites are present with high stoichiometry as initially reported 3, or low stoichiometry and rare 7 also remained to be resolved. It remained unclear why those studies produced divergent m 1A maps, and if m 1A exists at transcription-start sites or start codons, or neither, and why these particular sites are so prominent in m 1A-mapping studies. A re-analysis of these data showed that many of the sites that were mapped internally within the 5′UTR were actually transcription-start sites 9. Based on this location, it was proposed that m 1A forms a novel cap structure in which m 1A immediately follows the 7-methylguanosine (m 7G) cap of mRNA (m 7G-ppp-m 1A). In mRNAs, the majority of sites were found in the 5′UTR 22 of which the authors localized to the first nucleotide of the transcript. The second study mapped m 1A to 740 sites, 473 of which were in mRNA and lncRNA 8. Twelve other sites were detected at very low stoichiometry. Ultimately it was concluded that only two mRNAs contained high-confidence m 1A sites: C9orf100 and MT-ND5, a cytosolic and a mitochondrial mRNA, respectively 7. Thus it was concluded that mRNA fragments from the 5′UTR may be nonspecifically enriched during immunoprecipitation 7. Although mRNA fragments from 5′UTRs and start codon-proximal regions were immunoprecipitated, these fragments did not generate misincorporations. Using this approach, m 1A was rarely observed in the RNA immunoprecipitated with m 1A antibodies 7. In that study, the antibody-bound RNA was reverse transcribed with an enzyme that efficiently introduces misincorporations at m 1A. Subsequent work reported different distributions for m 1A, one arguing that m 1A was exceptionally rare in mRNA 7. Notably, most m 1A sites were located near start codons and proposed to provide a novel form of translational regulation 3. One study estimated the average stoichiometry of mapped m 1A sites at 20% 3. This antibody was previously shown to recognize m 1A-containing RNAs 6. Both studies mapped m 1A in thousands of mRNAs by sequencing mRNA fragments immunoprecipitated with a monoclonal antibody (clone AMA-2) commercially distributed by MBL Bioscience, which was originally raised against KLH-conjugated 1-methyladenosine 5. Two studies later identified N 1-methyladenosine (m 1A) as another abundant epitranscriptomic modification 3, 4. The initial concept of the epitranscriptome was born with the transcriptome-wide mapping of thousands of internally located modified nucleotide N 6-methyladenosine (m 6A) residues in the transcriptome 1, 2. These results demonstrate that high-stoichiometry m 1A sites are exceedingly rare in mRNAs and that previous mappings of m 1A to 5’UTRs were the result of antibody cross-reactivity to the 5’ cap. 
A different m 1A antibody that lacks cap-binding cross-reactivity does not show enriched binding in 5’UTRs. However, further analysis revealed that these were false-positives caused by binding of the antibody to the m 7G-cap. Using this approach, we detected appreciable levels of m 1A only in one mRNA: the mitochondrial MT-ND5 transcript. As an alternative approach, we also developed an antibody-based m 1A-mapping approach to detect m 1A at single-nucleotide resolution, and confirmed that the commonly used m 1A antibody maps sites to the transcription-start site in mRNA 5’UTRs. We developed a bioinformatic approach to discover m 1A and other modifications in mRNA throughout the transcriptome by analyzing preexisting ultra-deep RNA-Seq data for modification-induced misincorporations. N 1-methyladenosine (m 1A) was proposed to be a highly prevalent modification in mRNA 5’UTRs based on mapping studies using an m 1A-binding antibody.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
